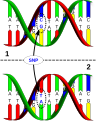
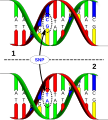
Construction process
This file was derived from a 3D model of DNA, converted to SVG and coloured using David Eccles' STL2SVG script:
type=orig; ~/scripts/stl2svg.pl ./DNA_linear_complete_helix1.stl:330 ./DNA_linear_complete_helix2.stl:200 ./DNA_linear_complete_${type}_A.stl:140 ./DNA_linear_complete_${type}_C.stl:250 ./DNA_linear_complete_${type}_G.stl:90 ./DNA_linear_complete_${type}_T.stl:30 > out_${type}.svg
type=mut; ~/scripts/stl2svg.pl ./DNA_linear_complete_helix1.stl:330 ./DNA_linear_complete_helix2.stl:200 ./DNA_linear_complete_${type}_A.stl:140 ./DNA_linear_complete_${type}_C.stl:250 ./DNA_linear_complete_${type}_G.stl:90 ./DNA_linear_complete_${type}_T.stl:30 > out_${type}.svg
The DNA models were then combined and annotated using Inkscape. The DNA backbone for the model is a pentagon extruded over a sine wave using David Eccles' guided path extrude script . The model source file (in OpenSCAD format) is shown below:
use <guided_extrude.scad>;
hl = 100; // helix length
hp = 33.2; // helix pitch [in angstroms]
hr = 10; // helix radius [in angstroms]
bbr = 1.5; // backbone radius
loops = hl / hp;
// random bases
//bases = rands(0, 4, ceil(360 * loops / 34.3),1);
// *GRINGENE* -- TAA GGN MGN ATH AAY GGN GAR AAY GAR TGA
// -- TAA GGC AGG ATC AAC GGC GAG AAC GAG TGA
// A = 0; G = 1; C = 2; T = 3
// [different from my usual order,
// to simplify the 3D model logic]
bases = [3,3,3, 1,1,2, 0,1,1, 0,3,2, 0,0,2,
1,1,2, 1,0,1, 0,0,2, 1,0,1, 3,1,0];
bAng = atan2(sin(120) - sin(0), cos(120) - cos(0));
drawMode = "all";
module lineTo(x1, x2){
hull(){
translate(x1) sphere(r=0.25, $fn=5);
translate(x2) sphere(r=0.25, $fn=5);
}
}
backbone_profile = [for(th = [0:72:359]) [bbr*cos(th),
bbr*sin(th)*1]];
inc = floor($t * 30);
thf = ($t * 30) - inc;
h1limit = (360 * loops);
h1jump = (360 * loops);
helix_1 = [for(th = [(thf*34.3):(34.3/2):h1jump])
[hr * cos(th), hr * sin(th), hl * th / (360 * loops)]];
helix_2 = [for(th = [120:(34.3/2):(360 * loops+120)])
[hr * cos(th), hr * sin(th), hl * (th-120) / (360 * loops)]];
module purine(){
linear_extrude(height=0.75, center=true){
// average hydrogen bond length in water: 1.97 A
// https://en.wiki.x.io/wiki/Hydrogen_bond#Structural_details
translate([-0.985,0])
// scale: average of C-C and C=C bond length
scale(1.435) translate([-2,0]) rotate(12) rotate(18){
rotate(-30) translate([1,0]) circle(r=1, $fn=6);
color("blue")
rotate(36) translate([-1 / (2*sin(36)),0])
circle(r=1 / (2*sin(36)), $fn=5);
}
}
}
module pyrimidine(){
linear_extrude(height=0.75, center=true){
// average hydrogen bond length in water: 1.97 A
// https://en.wiki.x.io/wiki/Hydrogen_bond#Structural_details
translate([-0.985,0])
scale(1.435) translate([-2, 0]) translate([1,0])
circle(r=1, $fn=6);
}
}
$vpt = [0, 0, 0];
//$vpr = [310, 105, 10];
$vpr = [0, 0, 0];
rotate([310, 105, 130]) translate([0,0,-hl/2]) {
if(drawMode == "all" || drawMode == "helix1") color("lightblue")
mapExtrude("vertCylinder", backbone_profile, helix_1);
if(drawMode == "all" || drawMode == "helix2") color("pink")
mapExtrude("vertCylinder", backbone_profile, helix_2);
for(thb = [inc:(360 * loops / 34.3 + inc)]) {
thi = thb-inc;
th = (thi-thf) * 34.3;
thisBase = bases[floor(thb%30)];
doPur = (thisBase < 2);
// base bond has a -1.2° angle;
// not quite sure how to implement that
baseFrac = (doPur ? 0.55 : 0.45);
baseFInv = 1 - baseFrac;
translate([0,0,hl * th / (360 * loops)]) rotate([-1.2,0,0]){
if(drawMode == "all" || drawMode == "helix2") color("pink")
lineTo([hr * cos(th)*(baseFrac-0.15) +
hr * cos(th+120) * (baseFrac+0.15),
hr * sin(th)*(baseFrac-0.15) +
hr * sin(th+120) * (baseFrac+0.15)],
[hr * cos(th+120), hr * sin(th+120)]);
if(th < (h1jump))
if(drawMode == "all" || drawMode == "helix1") color("lightblue")
lineTo([hr * cos(th), hr * sin(th)],
[hr * cos(th)*(baseFrac+0.15) +
hr * cos(th+120) * (baseFrac-0.15),
hr * sin(th)*(baseFrac+0.15) +
hr * sin(th+120) * (baseFrac-0.15)]);
if(drawMode == "all" ||
(drawMode == "A" && thisBase == 0) ||
(drawMode == "G" && thisBase == 1) ||
(drawMode == "C" && thisBase == 2) ||
(drawMode == "T" && thisBase == 3)
)
color((thisBase < 1) ? "green" :
(thisBase < 2) ? "gold" :
(thisBase < 3) ? "blue" :
"red")
translate([hr * cos(th)*baseFrac + hr * cos(th+120) * baseFInv,
hr * sin(th)*baseFrac + hr * sin(th+120) * baseFInv])
rotate(180 + bAng + th) if(doPur) {
purine(); } else { pyrimidine(); };
if(drawMode == "all" ||
(drawMode == "A" && thisBase == 3) ||
(drawMode == "G" && thisBase == 2) ||
(drawMode == "C" && thisBase == 1) ||
(drawMode == "T" && thisBase == 0)
)
if(th < (h1jump))
color((thisBase < 1) ? "red" :
(thisBase < 2) ? "blue" :
(thisBase < 3) ? "gold" :
"green")
translate([hr * cos(th)*baseFrac + hr * cos(th+120) * baseFInv,
hr * sin(th)*baseFrac + hr * sin(th+120) * baseFInv])
rotate(bAng+th) if(doPur) {
pyrimidine(); } else { purine(); };
}
}
if(drawMode == "all" || drawMode == "helix1") color("lightblue") {
translate(helix_1[len(helix_1)-1]) sphere(r=bbr, $fn=5);
translate(helix_1[0]) sphere(r=bbr, $fn=5);
}
if(drawMode == "all" || drawMode == "helix2") color("pink") {
translate(helix_2[0]) sphere(r=bbr, $fn=5);
translate(helix_2[len(helix_2)-1]) sphere(r=bbr, $fn=5);
}
}
この作品の著作権者である私は、この作品を以下のライセンスで提供します。
この文書は、フリーソフトウェア財団 発行のGNUフリー文書利用許諾書 (GNU Free Documentation License)GNUフリー文書利用許諾書 http://www.gnu.org/copyleft/fdl.html GFDL GNU Free Documentation License true true
帰属: SNP model by David Eccles (gringer)
あなたは以下の条件に従う場合に限り、自由に
共有 – 本作品を複製、頒布、展示、実演できます。再構成 – 二次的著作物を作成できます。
あなたの従うべき条件は以下の通りです。
表示 – あなたは適切なクレジットを表示し、ライセンスへのリンクを提供し、変更があったらその旨を示さなければなりません。これらは合理的であればどのような方法で行っても構いませんが、許諾者があなたやあなたの利用行為を支持していると示唆するような方法は除きます。 https://creativecommons.org/licenses/by/4.0 CC BY 4.0 Creative Commons Attribution 4.0 true true あなたは上記のライセンスから、どれか一つ以上を選択できます。
英語 DNA sequence variation in a population. A SNP is just a single nucleotide difference in the genome. The upper DNA molecule differs from the lower DNA molecule at a single base-pair location (a G/A polymorphism)